Research
RNA localization in homeostasis and disease
Targeting of RNA transcripts to diverse subcellular localizations has been observed in many different organisms and was identified as a general regulatory mechanism that controls gene expression on a transcriptome-wide scale. It has important functions in development and tissue homeostasis and attracts increasing interest for its role in cancer progression and metastasis. Yet, the mechanisms and RNA sequence elements that target most transcripts to specific subcellular localizations remain obscure. Most likely because cis-acting elements greatly vary in length and can act synergistically, it has been challenging to identify them based on primary sequence alone.
We employ single-molecule imaging techniques in combination with 3D organoid model systems to unravel the molecular mechanisms that target individual RNA transcripts to diverse subcellular localizations, cell types and tissues.
For more information, please visit our website (https://voigtlab.org)

Dynamics and mechanisms underlying RNA localization
Live single-molecule imaging of mRNA reporter transcripts in murine intestinal organoids to investigate how RNA localization regulates gene expression in a spatiotemporal context.
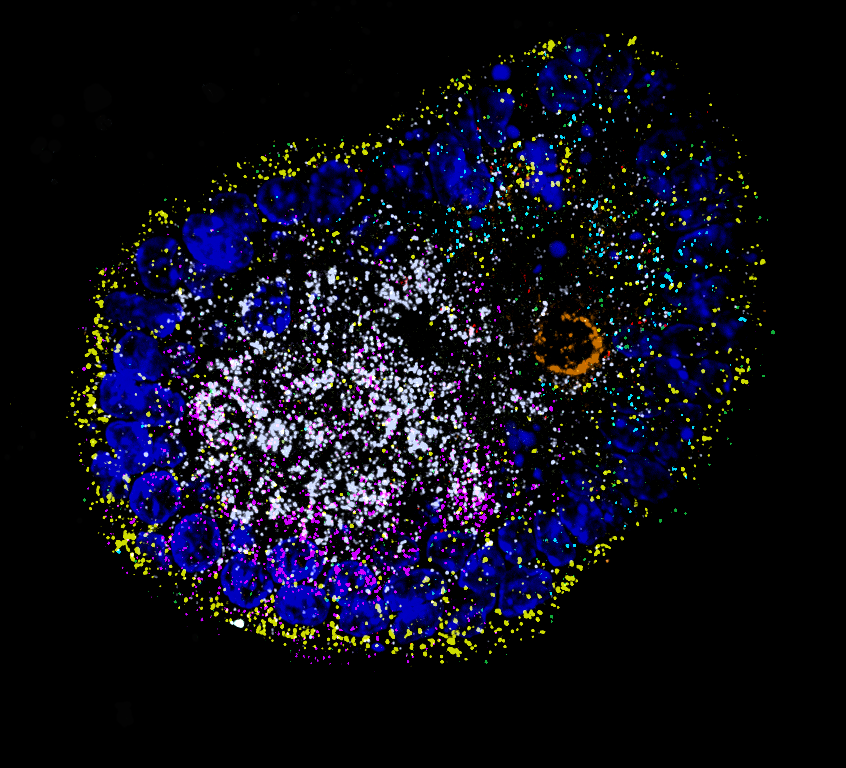
Global RNA localization patterns
Imaging-based spatial transcriptomics for parallel visualization of localization biases assumed by endogenous mRNA transcripts in different organoid models and physiological contexts.
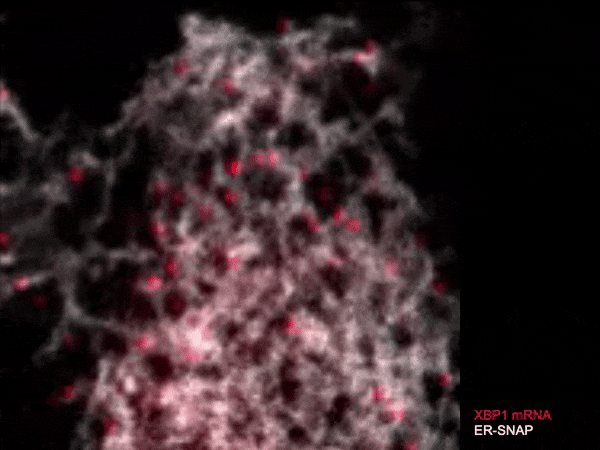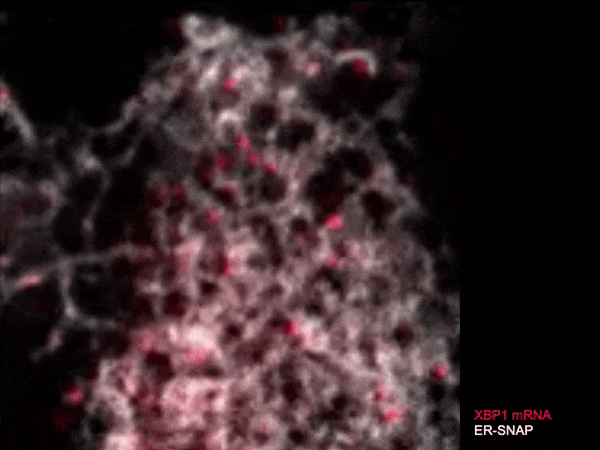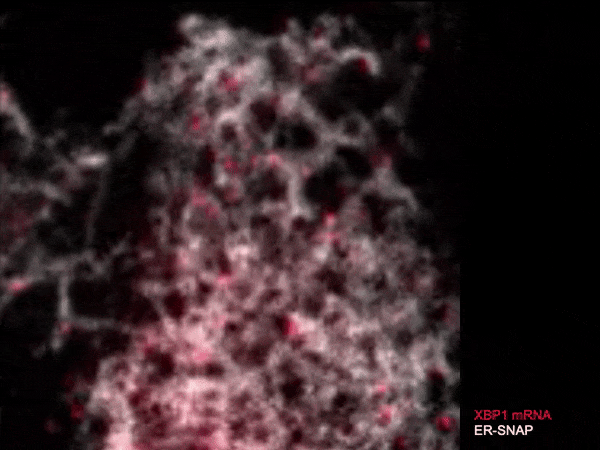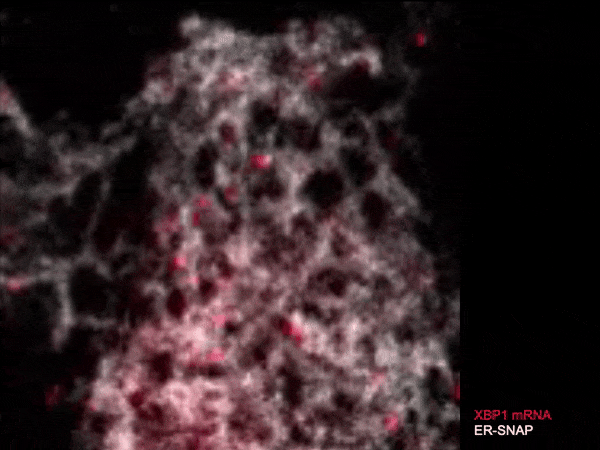
Live single-molecule imaging of XBP1 reporter mRNAs to investigate the dynamics of XBP1 recruitment to IRE1α oligomers in the ER membrane of individual HeLa cells.
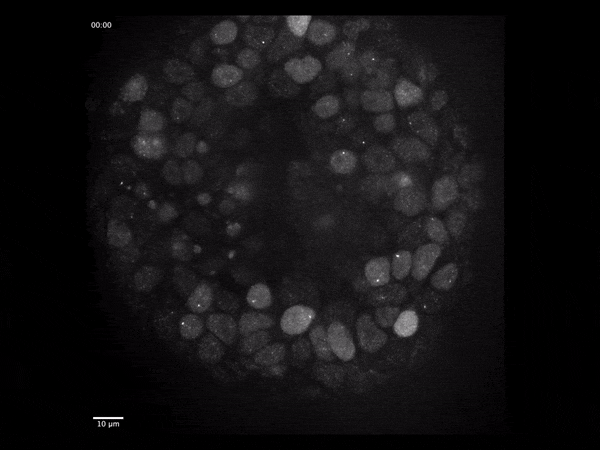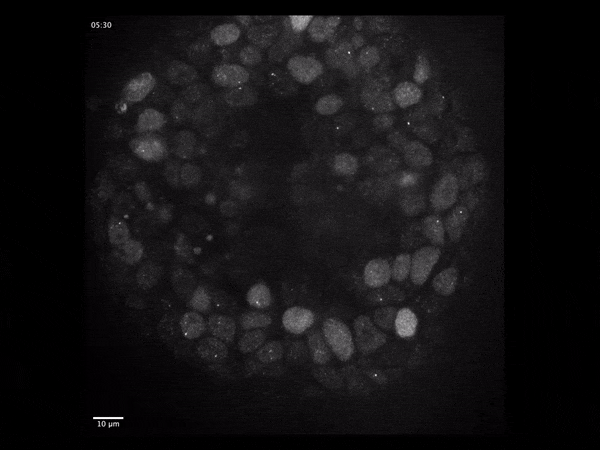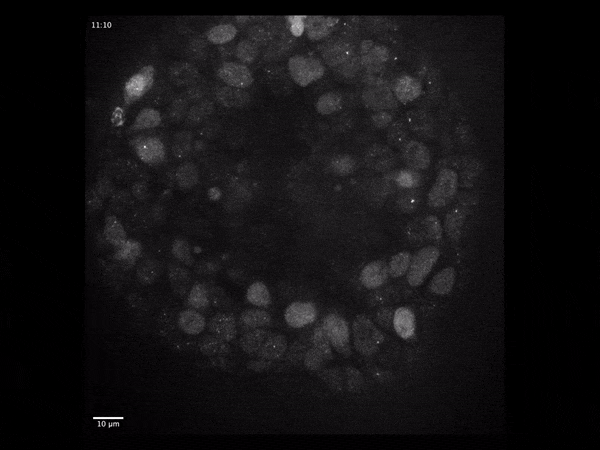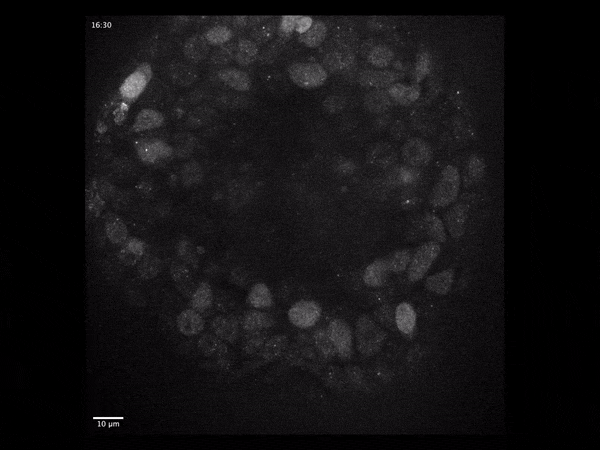
Gene expression dynamics during early human brain organoid development
Live single-molecule imaging of endogenously-tagged mRNA transcripts to quantify particle dynamics and transcriptional regulation using human iPSC-derived cerebral organoids.