Navigation auf uzh.ch
Navigation auf uzh.ch
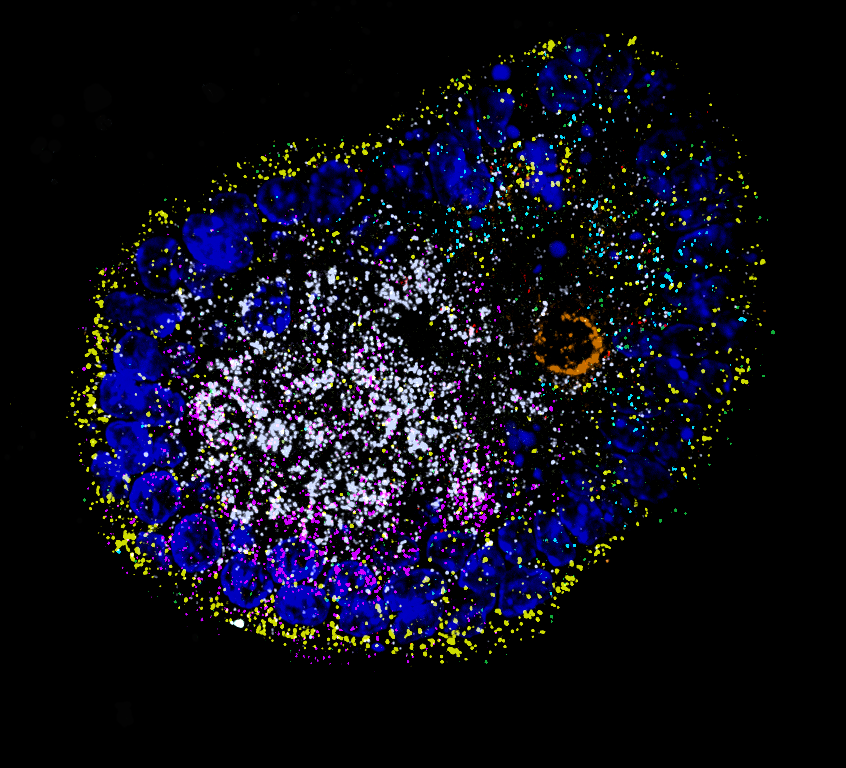
In order to map RNA localization patterns on a transcriptome-wide scale, we take a holistic and unbiased approach to identify novel cis-acting elements and correlate RNA localization with physiological function:
We use imaging-based spatial transcriptomics to systematically record localization patterns of many different endogenous transcripts in parallel and employ external stimuli to quantify changes in RNA localization biases, which will allow us to identify novel targeting motifs. For validation of our findings, we use single-cell genomics in collaboration with Treutlein lab, ETH Zurich DBSSE.
Our model systems are primarily murine intestinal organoids, which robustly recapitulate RNA localization biases that were previously described in literature. As well as more recently human iPSC-derived cerebral cysts and brain organoids that we have started to characterize as a multicellular model system to study RNA targeting in neuronal processes and at synapses.