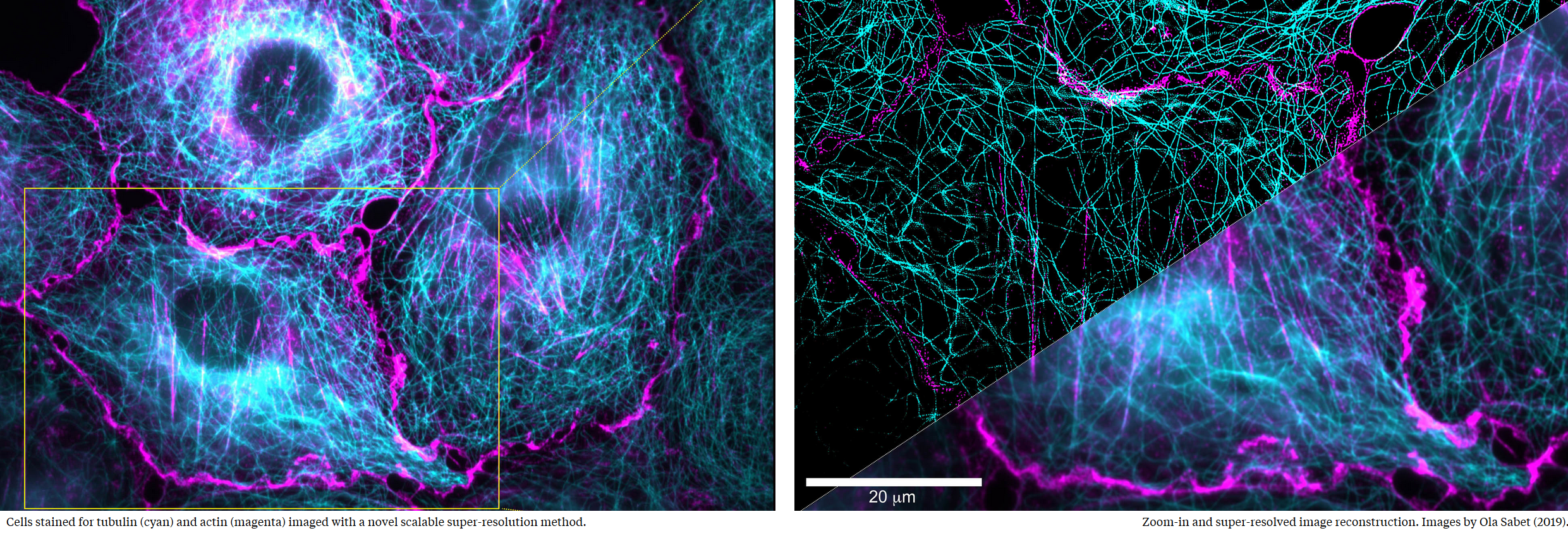
Crossing scales
In biology, many properties that are apparent at one scale cannot be reduced to properties of lower-scale components, yet they are determined by them.
One of the greatest scientific challenges of our era is to connect the different biological scales. We have reached a highly detailed understanding of life at individual scales, ranging from atomic structures of multi-subunit molecular machines to complex multi-species ecosystems, but we understand surprisingly little about how these scales are connected. The principles by which this is achieved may be scale-invariant, and rely on emergence and self-organisation, by which multiple local and often weak interactions between components at a lower scale give rise to new properties in form and function at a higher scale. The rules underlying these processes represent the grammar by which the book of life, the genome of an organism, is read. Our poor understanding of this grammar has become one of the biggest missing links in modern-day life science research, as the full potential of transformative technologies based on stem cells, genetic and tissue engineering, quantitative single-cell technologies, and large data sciences using multi-omics measurements will only be unlocked once we know these rules.
EMERGENCE AND SELF-ORGANISATION
Our group has pioneered research that specifically focuses on the rules by which biological scales are connected and we are dedicated to further dissect these emergent self-organising properties of living systems. We apply this research at multiple levels, to understand how individual gene products, mRNA transcripts and proteins, are patterned within cells, form compartments, and give rise to variation in gene expression. We study how the collective behaviour of individual organelles leads to cellular-scale functional properties, and how multiple cells together drive the emergence of different cellular states and spatial patterns of phenotypic variability. Moreover, we are starting to apply our scientific vision and technological expertise to the development of novel forms for post-genomics diagnostics, which aim at revealing scale-crossing effects in the emergence of disease phenotypes and responses to drugs.
SCALE-CROSSING TECHNOLOGIES
To study how properties at higher scales emerge from the collective behaviour of lower-scale interactions, and, importantly, how this feeds back on lower-scale behaviour, technologies are required that can collect measurements at multiple scales simultaneously. This can uncover previously hidden scale-crossing effects that offer crucial novel insights, such as realizing that the variability in a cell fate choice is not random, but strongly influenced by the population context of a cell, which emerges from the collective action of cells in a population. These insights can only be obtained when these properties are quantified for large numbers of individual cells in situ. One of the main approaches of the laboratory to collect such datasets is by means of imaging. To advance our capabilities to collect scale-crossing datasets, we develop new advanced microscopy methods, cellular computer vision algorithms, and apply the latest approaches in machine learning and information theory. One continuing quest in our lab is to extend the range of length and time units we can cross in such measurements, and to achieve this for as many different molecular species as possible. An essential property of the methods we develop is that they can scale to high-throughput automated applications in order to collect the large datasets necessary for revealing scale-crossing effects. This for instance implies that pushing into the realm of sub-diffraction limited imaging must be compatible with collecting these measurements across large surfaces in a reasonable time frame. Or, that time-lapse imaging of single molecules can be done in large numbers of single cells over long periods of time without photo-damaging the cells. This demand for scalability often requires innovative approaches.